m6A Detection via Nanopore
Intensive research focus in recent decades has been devoted to understanding the epigenetic and biomolecular landscape that underlie glioma. However, little attention has been given to modifications in mRNA that affect mRNA stability, nuclear export, splicing, and translation to protein. One such modification is N6-Methyladenosine (or "m6A"), a methyl group attached to mRNA at adenine nucleotides. Our lab is currently profiling patient glioma tissue to look for distinct differences in mRNA m6A modifications that impact tumor progression and growth.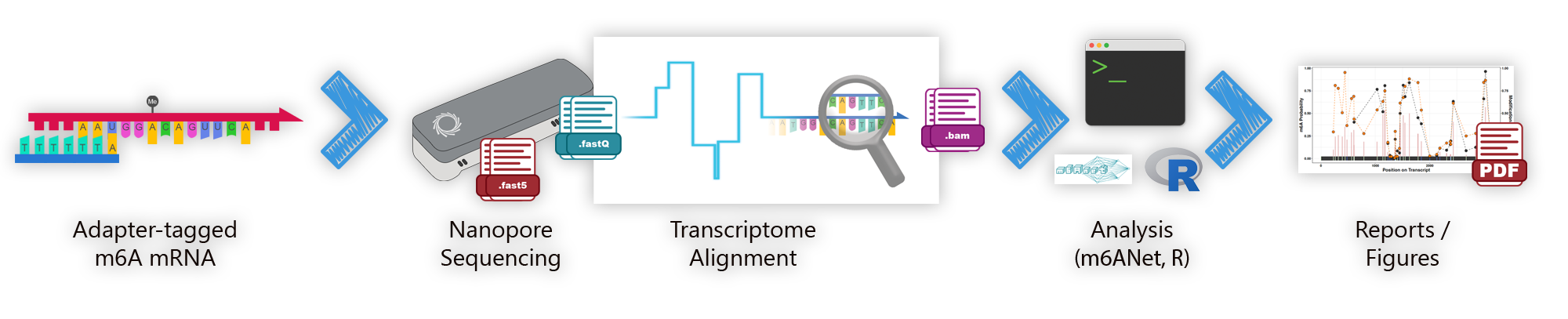
We currently achieve this by using direct RNA sequencing through the Oxford Nanopore Technology's minION system (shown above). RNA strands fed through nanoscopic pores within minION flow cells disrupt ionic currents in a base dependent manner (as well as upon modifications such as m6A). These signals can be determined to be modifications using sophisticated algorithms, such as EpiNano, Nanom6A, and m6ANet. Our goal in the lab is determine which modifications, and where, are critical for tumor development and progression.
